SDS-PAGE

SDS-PAGE, sodium dodecyl sulfate polyacrylamide gel electrophoresis, is a technique widely used in biochemistry, forensics, genetics and molecular biology to separate proteins according to their electrophoretic mobility (a function of length of polypeptide chain or molecular weight). SDS gel electrophoresis of samples having identical charge per unit mass due to binding of SDS results in fractionation by size.
Contents |
Procedure
Tissue preparation
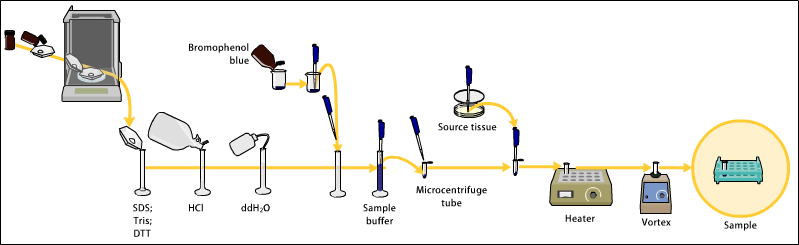
Samples may be taken from whole tissue or from cell culture. In most cases, solid tissues are first broken down mechanically using a blender (for larger sample volumes), using a homogenizer (smaller volumes), or by sonication. Cells may also be broken open by one of the above mechanical methods. However, it should be noted that bacteria, virus or environmental samples can be the source of protein and thus Western blotting is not restricted to cellular studies only.
A combination of biochemical and mechanical techniques – including various types of filtration and centrifugation – can be used to separate different cell compartments and organelles.
The solution of proteins to be analyzed is mixed with SDS, an anionic detergent which denatures secondary and non–disulfide–linked tertiary structures, and applies a negative charge to each protein in proportion to its mass. Heating the samples to at least 60 degrees C shakes up the molecules, helping SDS to bind. [1] [2] [3] [4]
A tracking dye may be added to the protein solution (of a size smaller than protein) to allow the experimenter to track the progress of the protein solution through the gel during the electrophoretic run.
Preparing acrylamide gels
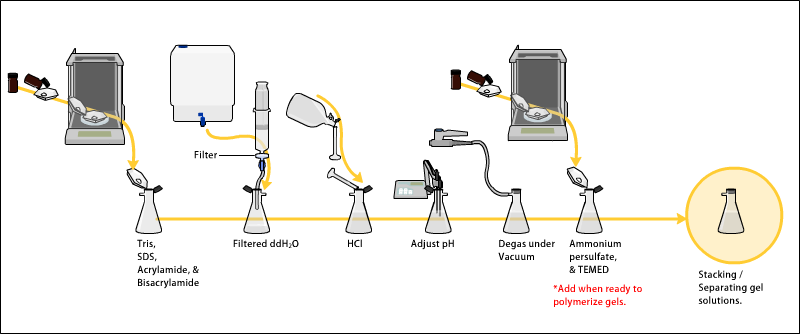
The gels generally consist of acrylamide, bisacrylamide, SDS, and a Tris-Cl buffer with adjusted pH. The solution is degassed under a vacuum to prevent air bubbles during polymerization. [5] Ammonium persulfate and TEMED are added when the gel is ready to be polymerized. The separating or resolving gel is usually more basic and has a higher polyacrylamide content than the loading gel. [6]
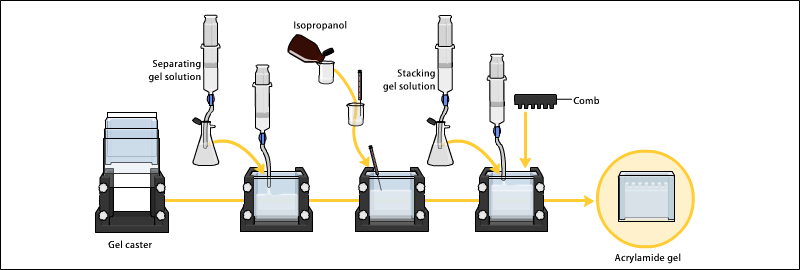
Gels are polymerized in a gel caster. First the separating gel is poured and allowed to polymerize. Next a thin layer of isopropanol is added. Next the loading gel is poured and a comb is placed to create the wells. After the loading gel is polymerized the comb can be removed and the gel is ready for electrophoresis.
Electrophoresis
First the anode and cathode buffers are prepared. The anode buffer usually contains Tris-Cl, distilled deionized water and is adjusted to a higher pH than the cathode buffer. The cathode buffer contains SDS, Tris, Tricine, and distilled deionized water. [7] [8]

The electrophoresis apparatus is set up with cathode buffer covering the gel in the negative electrode chamber, and anode buffer in the lower positive electrode chamber. Next, the denatured sample proteins are added to the wells one end of the gel with a syringe or pipette. Finally, the apparatus is hooked up to a power source under appropriate running conditions to separate the protein bands.
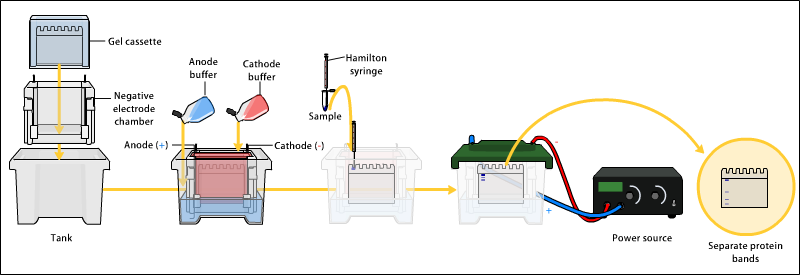
An electric field is applied across the gel, causing the negatively-charged proteins to migrate across the gel towards the positive (+) electrode (anode). Depending on their size, each protein will move differently through the gel matrix: short proteins will more easily fit through the pores in the gel, while larger ones will have more difficulty (they encounter more resistance). After a set amount of time (usually a few hours- though this depends on the voltage applied across the gel; higher voltages run faster but tend to produce somewhat poorer resolution), the proteins will have differentially migrated based on their size; smaller proteins will have traveled farther down the gel, while larger ones will have remained closer to the point of origin. Therefore, proteins may be separated roughly according to size (and therefore, molecular weight), certain glycoproteins behave anomalously on SDS gels.
Staining

Following electrophoresis, the gel may be stained (most commonly with Coomassie Brilliant Blue R-250 or silver stain), allowing visualization of the separated proteins, or processed further (e.g. Western blot). After staining, different proteins will appear as distinct bands within the gel. It is common to run molecular weight size markers of known molecular weight in a separate lane in the gel, in order to calibrate the gel and determine the weight of unknown proteins by comparing the distance traveled relative to the marker. The gel is actually formed because the acrylamide solution contains a small amount, generally about 1 part in 35 of bisacrylamide, which can form cross-links between two polyacrylamide molecules. The ratio of acrylamide to bisacrylamide can be varied for special purposes. The acrylamide concentration of the gel can also be varied, generally in the range from 5% to 25%. Lower percentage gels are better for resolving very high molecular weight proteins, while much higher percentages are needed to resolve smaller proteins. Determining how much of the various solutions to mix together to make gels of particular acrylamide concentration can be done on line
Gel electrophoresis is usually the first choice as an assay of protein purity due to its reliability and ease. The presence of SDS and the denaturing step causes proteins to be separated solely based on size. False negatives and positives are possible. A comigrating contaminant can appear as the same band as the desired protein. This comigration could also cause a protein to run at a different position or to not be able to penetrate the gel. This is why it is important to stain the entire gel including the stacking section. Coomassie Brilliant Blue will also bind with less affinity to glycoproteins and fibrous proteins, which interferes with quantification.
Chemical ingredients and their roles
Polyacrylamide gel (PAG) had been known as a potential embedding medium for sectioning tissues as early as 1964. Two independent groups, Davis and Raymond, employed PAG in electrophoresis in 1959.[9] [10] It possesses several electrophoretically desirable features that make it a versatile medium. PAGE separates protein molecules according to both size and charge. It is a synthetic gel, thermo-stable, transparent, strong, relatively chemically inert, can be prepared with a wide range of average pore sizes [11]. The pore size of a gel is determined by two factors, the total amount of acrylamide present (%T) (T = Total acrylamide-bisacrylamide monomer concentration) and the amount of cross-linker (%C) (C = Crosslinker concentration). Pore size decreases with increasing %T; with cross-linking, 5%C gives the smallest pore size. Any increase or decrease in %C increases the pore size, as pore size with respect to %C is a parabolic function with vertex as 5%C. This appears to be because of nonhomogeneous bundling of strands in the gel.
This gel material can also withstand high voltage gradients, feasible for various staining and destaining procedures, and can be digested to extract separated fractions or dried for autoradiography and permanent recording. DISC electrophoresis utilizes gels of different pore sizes. [12] [13] The name DISC was derived from the discontinuities in the electrophoretic matrix and coincidentally from the discoid shape of the separated zones of ions. There are two layers of gel, namely stacking or spacer gel, and resolving or separating gel.

Stacking gel
The stacking gel is a large pore PAG (4%T). This gel is prepared with Tris/HCl buffer pH 6.8 of about 2 pH units lower than that of electrophoresis buffer (Tris/Glycine). These conditions provide an environment for Kohlrausch reactions determining molar conductivity, as a result, SDS-coated proteins are concentrated to several fold and a thin starting zone of the order of 19 μm is achieved in a few minutes. This gel is cast over the resolving gel. The height of the stacking gel region is always maintained more than double the height and the volume of the sample to be applied. This is based on isotachophoresis.
Chemical ingredients
- Tris (tris (hydroxy methyl) aminomethane) (C4H11NO3; mW: 121.14). It has been used as a buffer because it is an innocuous substance to most proteins. Its pKa is 8.3 at 20 °C, making it a very satisfactory buffer in the pH range from roughly 7 to 9.
- Glycine (Amino Acetic Acid) (C2H5NO2; mW: 75.07). Glycine has been used as the source of trailing ion or slow ion because its pKa is 9.69 and mobility of glycinate are such that the effective mobility can be set at a value below that of the slowest known proteins of net negative charge in the pH range. The minimum pH of this range is approximately 8.0.
- Acrylamide (C3H5NO; mW: 71.08). It is a white crystalline powder. While dissolving in water, autopolymerization of acrylamide takes place. It is a slow spontaneous process by which acrylamide molecules join together by head on tail fashion. But in presence of free radicals generating system, acrylamide monomers are activated into a free-radical state. These activated monomers polymerise quickly and form long chain polymers. This kind of reaction is known as Vinyl addition polymerisation. A solution of these polymer chains becomes viscous but does not form a gel, because the chains simply slide over one another. Gel formation requires hooking various chains together. Acrylamide is a neurotoxin. It is also essential to store acrylamide in a cool dark and dry place to reduce autopolymerisation and hydrolysis.
- Bisacrylamide (N,N'-Methylenebisacrylamide) (C7H10N2O2; mW: 154.17). Bisacrylamide is the most frequently used cross linking agent for poly acrylamide gels. Chemically it is thought of having two-acrylamide molecules coupled head to head at their non-reactive ends.
- Sodium Dodecyl Sulfate (SDS) (C12H25NaO4S; mW: 288.38). SDS is the most common dissociating agent used to denature native proteins to individual polypeptides. When a protein mixture is heated to 100 °C in presence of SDS, the detergent wraps around the polypeptide backbone. It binds to polypeptides in a constant weight ratio of 1.4 g/g of polypeptide. In this process, the intrinsic charges of polypeptides becomes negligible when compared to the negative charges contributed by SDS. Thus polypeptides after treatment becomes a rod like structure possessing a uniform charge density, that is same net negative charge per unit length. Mobilities of these proteins will be a linear function of the logarithms of their molecular weights.
- Without SDS, different proteins with similar molecular weights would migrate differently due to differences in mass charge ratio, as each protein has an isoelectric point and molecular weight particular to its primary structure. This is known as Native PAGE. Adding SDS solves this problem, as it binds to and unfolds the protein, giving a near uniform negative charge along the length of the polypeptide.
- Ammonium persulfate (APS) (N2H8S2O8; mW: 228.2). APS is an initiator for gel formation.
- TEMED (N, N, N', N'-tetramethylethylenediamine) (C6H16N2; mW: 116.21). Chemical polymerisation of acrylamide gel is used for SDS-PAGE. It can be initiated by ammonium persulfate and the quaternary amine, N,N,N',N'-tetramethylethylenediamine (TEMED). The rate of polymerisation and the properties of the resulting gel depends on the concentration of APS and TEMED. Increasing the amount of APS and TEMED results in a decrease in the average polymer chain length, an increase in gel turbidity and a decrease in gel elasticity. Decreasing the amount of initiators shows the reverse effect. The lowest catalytic concentrations that will allow polymerisation in the optimal period of time should be used. APS and TEMED are used, approximately in equimolar concentrations in the range of 1 to 10 mM.
Chemicals for processing and visualization
The following chemicals are used for processing of the gel and the protein samples visualized in it:
- Bromophenol blue (BPB) (3',3",5',5" tetrabromophenolsulfonphthalein) (C19H10Br4O5S; mW: 669.99). BPB is the universal marker dye. Proteins and nucleic acids are mostly colourless. When they are subjected to electrophoresis, it is important to stop the run before they run off the gel. BPB is the most commonly employed tracking dye, because it is viable in alkali and neutral pH, it is a small molecule, it is ionisable and it is negatively charged above pH 4.6 and hence moves towards the anode. Being a small molecule it moves ahead of most proteins and nucleic acids. As it reaches the anodic end of the electrophoresis medium electrophoresis is stopped. It can bind with proteins weakly and give blue colour.
- Glycerol (C3H8O3; mW: 92.09). It is a preservative and a weighing agent. Addition of glycerol (20-30 or 50%) is often recommended for the storage of enzymes. Glycerol maintains the protein solution at very low temperature, without freezing. It also helps to weigh down the sample into the wells without being spread while loading.
- Coomassie Brilliant Blue R-250 (CBB)(C45H44N3NaO7S2; mW: 825.97). CBB is the most popular protein stain. It is an anionic dye, which binds with proteins non-specifically. The structure of CBB is predominantly non-polar. So is usually used (0.025%) in methanolic solution (40%) and acetic acid (7%). Proteins in the gel are fixed by acetic acid and simultaneously stained. The excess dye incorporated in the gel can be removed by destaining with the same solution but without the dye. The proteins are detected as blue bands on a clear background. As SDS is also anionic, it may interfere with staining process. Therefore, large volume of staining solution is recommended, approximately ten times the volume of the gel.
- n-Butanol (C4H10O; mW: 74.12). Water saturated butanol is used as an overlay solution on the resolving gel.
- Dithiothreitol (DTT; C4H10O2S2; mW: 154.25). DTT is a reducing agent used to disrupt disulfide bonds to ensure the protein is fully denatured before loading on the gel; ensuring the protein runs uniformly. Traditionally the toxic and less potent 2-mercaptoethanol was used.
Reducing SDS-PAGE
Besides the addition of SDS, proteins may optionally be briefly heated to near boiling in the presence of a reducing agent, such as dithiothreitol (DTT) or traditionally 2-mercaptoethanol (beta-mercaptoethanol/BME), which further denatures the proteins by reducing disulfide linkages, thus overcoming some forms of tertiary protein folding, and breaking up quaternary protein structure (oligomeric subunits). This is known as reducing SDS-PAGE, and is most commonly used. Non-reducing SDS-PAGE (no boiling and no reducing agent) may be used when native structure is important in further analysis (e.g. enzyme activity, shown by the use of zymograms). For example, quantitative preparative native continuous polyacrylamide gel electrophoresis (QPNC-PAGE) is a new method for separating native metalloproteins in complex biological matrices.
Silver staining

In the 14th century the silver staining technique was developed for colouring the surface of glass. It has been used extensively for this purpose since the 16th century. The colour produced by the early silver stains ranged between light yellow and an orange-red. Camillo Golgi perfected the silver staining for the study of the nervous system. Golgi's method stains a limited number of cells at random in their entirety.[14] The exact chemical mechanism by which this happens is still largely unknown.[15] Silver staining was introduced by Kerenyi and Gallyas as a sensitive procedure to detect trace amounts of proteins in gels.[16] The technique has been extended to the study of other biological macromolecules that have been separated in a variety of supports.[17] Classical Coomassie Brilliant Blue staining can usually detect a 50 ng protein band, Silver staining increases the sensitivity typically 50 times. Many variables can influence the colour intensity and every protein has its own staining characteristics; clean glassware, pure reagents and water of highest purity are the key points to successful staining.[18]
Buffer systems
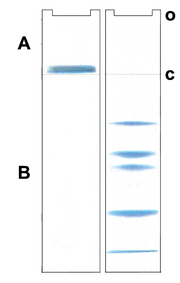

Most protein separations are performed using a "discontinuous" buffer system that significantly enhances the sharpness of the bands within the gel. During electrophoresis in a discontinuous gel system, an ion gradient is formed in the early stage of electrophoresis that causes all of the proteins to focus into a single sharp band. This occurs in a region of the gel that has larger pores so that the gel matrix does not retard the migration during the focusing or "stacking" event. Negative ions from the buffer in the tank then "outrun" the SDS-covered protein "stack" and eliminate the ion gradient so that the proteins subsequently separate by the sieving action in the lower, "resolving" region of the gel.
Many people continue to use a tris-glycine or "Laemmli" buffering system that stacks at a pH of 6.8 and resolves at a pH of ~8.3-9.0. These pHs promote disulfide bond formation between cysteine residues in the proteins, especially when they are present at high concentrations because the pKa of cysteine ranges from 8-9 and because reducing agent present in the loading buffer doesn't co-migrate with the proteins. Recent advances in buffering technology alleviate this problem by resolving the proteins at a pH well below the pKa of cysteine (e.g., bis-tris, pH 6.5) and include reducing agents (e.g. sodium bisulfite) that move into the gel ahead of the proteins to maintain a reducing environment. An additional benefit of using buffers with lower pHs is that the acrylamide gel is more stable so the gels can be stored for long periods of time before use.[19] [20]
SDS gradient gel electrophoresis of proteins

As voltage is applied, the anions (and negatively charged sample molecules) migrate toward the positive electrode (anode) in the lower chamber, the leading ion is Cl¯ ( high mobility and high concentration); glycinate is the trailing ion (low mobility and low concentration). SDS-protein particles do not migrate freely at the border between the Cl¯ of the gel buffer and the Gly¯ of the cathode buffer. Friedrich Kohlrausch found that Ohm's law also applies to dissolved electrolytes. Because of the voltage drop between the Cl- and Glycine-buffers, proteins are compressed (stacked) into micrometer thin layers. [21] The boundary moves through a pore gradient and the protein stack gradually disperses due to an frictional resistance increase of the gel matrix. Stacking and unstacking occurs continuously in the gradient gel, for every protein at a different position. For a complete protein unstacking the polyacrylamide-gel concentration must exceed 16% T. The two-gel system of "Laemmli" is a simple gradient gel. The pH discontinuity of the buffers is of no significance for the separation quality, and a "stacking-gel" with a different pH is not needed.
See also
- Capillary electrophoresis
- DNA electrophoresis
- Eastern blotting
- Electroblotting
- Electrophoresis
- Gel electrophoresis
- History of electrophoresis
- Isoelectric focusing
- Isotachophoresis
- Native Gel Electrophoresis
- Northern blotting
- Protein electrophoresis
- Southern blotting
- Two dimensional SDS-PAGE
- Western blotting
- Zymography
References
- ↑ Shapiro AL, Viñuela E, Maizel JV Jr. (September 1967). "Molecular weight estimation of polypeptide chains by electrophoresis in SDS-polyacrylamide gels.". Biochem Biophys Res Commun. 28 (5): 815–820. doi:10.1016/0006-291X(67)90391-9. PMID 4861258.
- ↑ Weber K, Osborn M (August 1969). "The reliability of molecular weight determinations by dodecyl sulfate-polyacrylamide gel electrophoresis.". J Biol Chem. 244 (16): 4406–4412. PMID 5806584.
- ↑ Laemmli UK (August 1970). "Cleavage of structural proteins during the assembly of the head of bacteriophage T4". Nature 227 (5259): 680–685. doi:10.1038/227680a0. PMID 5432063.
- ↑ Caprette, David. "SDS-PAGE". http://www.ruf.rice.edu/~bioslabs/studies/sds-page/denature.html. Retrieved 27 Sept 2009.
- ↑ "What is the meaning of de -gas the acrylamide gel mix?". http://www.protocol-online.org/biology-forums/posts/17740.html. Retrieved 28 Sept 2009.
- ↑ "SDS-PAGE". http://www.science.smith.edu/departments/Biochem/Biochem_353/sdspage.html. Retrieved 12 Sept 2009.
- ↑ Schägger, H; Von Jagow, G (1987). "Tricine-sodium dodecyl sulfate-polyacrylamide gel electrophoresis for the separation of proteins in the range from 1 to 100 kDa.". Anal. Biochem. 166 (2): 368–379. doi:10.1016/0003-2697(87)90587-2. PMID 2449095.
- ↑ Andrews. "SDS-PAGE". http://www.dwalab.ca/labman/recipes/PreparingtoRunTricineSDSGels.html. Retrieved 27 Sept 2009.
- ↑ Davis BJ, Ornstein L (1959). "A new high resolution electrophoresis method.". Delivered at the Society for the Study of Blood at the New York Academy of Medicine.
- ↑ Raymond S, Weintraub L. (1959). "Acrylamide gel as a supporting medium for zone electrophoresis.". Science 130: 711. doi:10.1126/science.130.3377.711. PMID 14436634.
- ↑ Rüchel R, Steere RL, Erbe EF (1978). "Transmission-electron microscopic observations of freeze-etched polyacrylamide gels.". J Chromatogr. 166: 563–575. doi:10.1016/S0021-9673(00)95641-3.
- ↑ Ornstein L (December 1964). "DISC ELECTROPHORESIS. I. BACKGROUND AND THEORY.". Ann N Y Acad Sci. 121: 321–349. doi:10.1111/j.1749-6632.1964.tb14207.x. PMID 14240533.
- ↑ Davis BJ (December 1964). "Disc Electrophoresis. 2, Method and application to human serum proteins". Ann. New York Acad. Sci 121: 404–427. doi:10.1111/j.1749-6632.1964.tb14213.x. PMID 14240539.
- ↑ Grant G (Oct 2007). "How the 1906 Nobel Prize in Physiology or Medicine was shared between Golgi and Cajal". Brain Res Rev 55 (2): 490–498. doi:10.1016/j.brainresrev.2006.11.004. PMID 17306375.
- ↑ Golgi C (1873). "Sulla struttura della sostanza grigia del cervello.". Gazzetta Medica Italiana (Lombardia) 33: 244–246.
- ↑ Kerenyi L, Gallyas F (1973). "Über Probleme der quantitiven Auswertung der mit physikalischer Entwicklung versilberten Agarelektrophoretogramme". Clin. Chim. Acta 47 (3): 425–436. doi:10.1016/0009-8981(73)90276-3. PMID 4744834.
- ↑ Switzer RC 3rd, Merril CR, Shifrin S (Sep 1979). "A highly sensitive silver stain for detecting proteins and peptides in polyacrylamide gels.". Anal Biochem. 98 (1): 231–237. doi:10.1016/0003-2697(79)90732-2. PMID 94518.
- ↑ Hempelmann E, Schulze M, Götze O (1984). "Free SH-groups are important for the polychromatic staining of proteins with silver nitrat". Neuhof V (ed)Electrophoresis '84 , Verlag Chemie Weinheim 1984: 328–330.
- ↑ Schägger H, von Jagow G (1987). "Tricine-sodium dodecyl sulfate-polyacrylamide gel electrophoresis for the separation of proteins in the range from 1 to 100 kDa.". Anal Biochem. 166 (2): 368–379. doi:10.1016/0003-2697(87)90587-2. PMID 2449095.
- ↑ Wiltfang J, Arold N, Neuhoff V (1991). "A new multiphasic buffer system for sodium dodecyl sulfate-polyacrylamide gel electrophoresis of proteins and peptides with molecular masses 100,000-1000, and their detection with picomolar sensitivity.". Electrophoresis 12 (5): 352–366. doi:10.1002/elps.1150120507. PMID 1718736.
- ↑ Kohlrausch F (1897). "Ueber Concentrations-Verschiebungen durch Electrolyse im Inneren von Lösungen und Lösungsgemischen.". Ann.J.Phys.u.Chem. 62: 209–239. doi:10.1002/andp.18972981002.
External links
- Demystifying SDS-PAGE Video
- Demystifying SDS-PAGE
- SDS-PAGE Calculator for customised recipes for TRIS Urea gels.
- 2-Dimensional Protein Gelelectrophoresis
- [1] Hempelmann E. SDS-Protein PAGE and Proteindetection by Silverstaining and Immunoblotting of Plasmodium falciparum proteins. in: Moll K, Ljungström J, Perlmann H, Scherf A, Wahlgren M (eds) Methods in Malaria Research, 5th edition, 2008, 263-266
|
|||||||||||||||||